I am still quite confused. As described in my posts above, I have deleted the page links in the bibliographic records created automatically during import from Zotero. This way I replaced [[Abstract]] with Abstract, for example. But I also removed the links from the names of the authors as listed within the authors property. Finally, I manually deleted pages corresponding to the authors. I did this step within Logseq. So far so good.
Now after switching between two synchronized computers I did refresh and reindexing and looking into the full list of pages in Logseq, the pages corresponding to authors are back there.
At first I thought that this must be some sync issue (I am actually struggling with syncing too), but I checked the pages corresponding to the imported publications and they seem to stay stripped of the links as I wanted. So I removed the pages generated for the authors, but after a while they are back again. As viewed in Logseq.
Apparently I am still missing some understanding of Logseq. If I now look into the corresponding directory/folder on my computer where Logseq data reside, there are no files corresponding to the authors. But within Logseq the All pages gives me a list that does contain the authors.
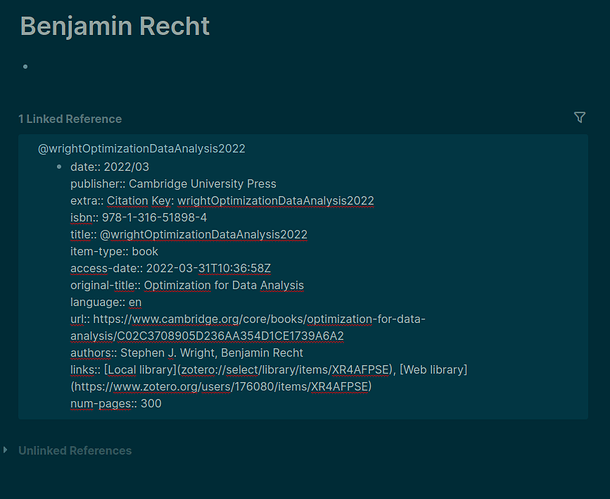
In fact, in this list all the authors’ pages have at least 1 back link. And these seem to go from the publication record. But now comes the (another) confusing part. See the two screenshots below.
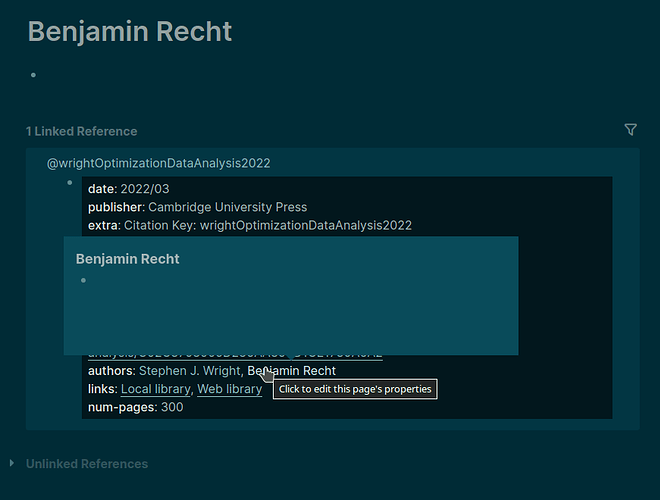
In the editing mode it is obvious that the authors property in the publication record is stripped of the double square brackets [[ ]]. Good. But as soon as I switch to the display mode (or whatever it is called) just by clicking elsewhere, the authors are displayed “clickable”.
How come?
To summarize, I have two questions:
- How come that in the list of pages in Logseq (clicking
All pages), some pages are displayed but no corresponding files are created in the directory (as viewed using a system browser). - How come that some block contains a property
authorsfollowed by a simple unlinked text, for exampleauthors:: Stephen J. Wright, Benjamin Recht, and yet their names are clickable (and corresponding author pages are listed as in the question 1)?